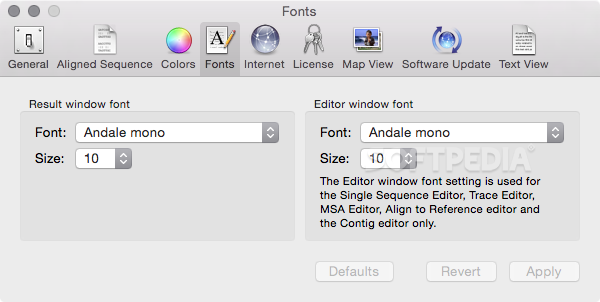
Getting Started with MacVector: An overview of primer design workflows in MacVector.Melissa Caimano on HOW DO I video guides to common molecular biology workflows.admin on HOW DO I video guides to common molecular biology workflows.mariam abdelmalak on Major release details – Summary.Brian on Designing primers and documenting In-Fusion Cloning with MacVector.Open up Automator.app, create a new Service. Hold Cmd and drag to invert the selection: If you don't want to use the mouse, an alternative way would be to use AppleScript instead. Primer sequences were as follows: reverse transcriptase. Chris on Designing primers and documenting In-Fusion Cloning with MacVector Select the files any view should work then move with the mouse cursor to the empty space next to the file list. Primer design was facilitated with MacVector Pro software version 16 (MacVector, Inc.How to call heterozygotes in trace files or Assembly Projects.Some alignment programs are MacVector (Oxford Molecular Ltd, Oxford, U.K.). MacVectorTip: How to Customize the Toolbars of MacVector windows In some cases, the auxotrophic marker gene is selected from an argB gene.MacVectorTip: Selecting the sequence from a single restriction enzyme site to the end of a linear sequence.Sequence Assembly: What can Assembler do for my lab?.If your sequence has multiple SNPs/genotypes/repeats, you can always then choose the right-click Delete Selected Reads option to remove those reads and start again on another set. Now you have the mate-pairs of the SNP reads selected and you can save all the selected reads using the right-click Export Selected Reads as FastA/Q option. Finally, right-click and choose Select Matching Pairs. This selects every read that aligns at that location with the G at that position. Then right-click (-click) and choose Select Overlapping Reads Containing Selected Sequence from the context sensitive menu. This helps ignore the occasional “bad” sequence, though, for most purposes, you can just select the one residue. To select all of the reads that contain the SNP, first select a few residues around that SNP, as shown above. Note that the Dots toolbar button is toggled on to help emphasize the mismatches Here we see an Align to Reference where about half the reads have obvious SNPs compared to the reference. 
NOTE: if your reference represents just a subset of the data in the NGS files, you might want to first filter the data using Align to Folder Now click the Add Seqs button and select and add your NGS data files. 
Take a reference sequence and choose Analyze->Align to Reference. When analyzing/assembling/aligning NGS data, there are many scenarios where you might want to separate out the reads representing different genotypes or variant sequences.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
